Research Lab
MolMicro of Bacterial Pathogens

29681
Lab Members
Lab leader
Senior Researcher
PhD students
Master Students
Research Interests
The bacterial cell wall is a complex structure which suffers constant synthesis and degradation to allow for growth and division. Besides its structural role, it also provides a communication platform with the environment.
In our lab, we study the cell wall of the human pathogen Staphylococcus aureus, leading cause of hospital-acquired infections, mainly due to its capacity to resist antibiotics. We combine biochemical and genetic approaches to understand the molecular mechanisms which associate the cell wall to the physiology and mechanisms of resistance of this pathogen. We have a specific interest in the study of biofilm formation, host interactions and application of biophysical methods.
Research Highlights
Finding new strategies to fight bacterial pathogens
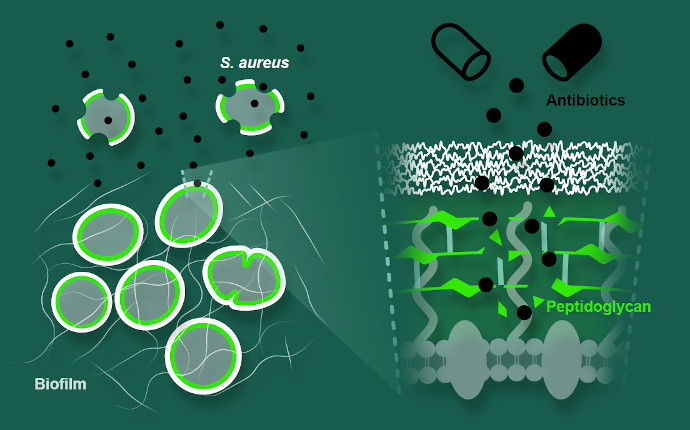
The number of antibiotics that remain efficient against bacterial pathogens, namely multidrug resistant strains, is alarmingly decreasing and new strategies are urgently needed to deal with this global crisis. One of our goals is to study cell wall weak spots that can be used in antibacterial strategies, such as peptidoglycan amidation. Also, we are characterizing a new two-peptide system from S. aureus that is involved in interspecies interactions. Furthermore, cork extractives are being explored for their antibacterial activity not only against human pathogens, S. aureus, E. faecalis. and E. coli, but also against prominent plant pathogens, namely Xyllela fastidiosa.
Potential of the Mediterranean Diet to increase quality of life – DM4Y
DM4Y project aims to explore the nutraceutical effects of plant based foods, namely of the Mediterranean diet, rich in plant-based foods, such as fruits, vegetables, and legumes, that has been associated with a reduction in overall mortality and multiple health benefits.
We will evaluate the impact of soup and fruit on cellular aging and inflammatory response at the blood plasma protein level.
Representative Projects
- “StaphOUT- Fighting Staphylococcus aureus - Peptidoglycan amidation as a new target”. FCT-MCTES. Total and Unit funding: €239,900. Rita Sobral (PI)
- “Microfluidics Liquid Crystal Based Bifunctional Bacterial Infection Sensor”, FCT-MCTES, Funding: €159,912, Rita Sobral (Co-PI)
- “Amidation of the peptidoglycan of Gram-positive bacteria: an unexplored potential target for antibiotics”, FCT-MCTES, Total funding: €165,000, Unit funding: €60,872, Rita Sobral (Collaborator)
- “A new target inside an old molecule: glutamate amidation of peptidoglycan”, ESCMID, Funding: €20,000, Rita Sobral (PI)
Selected Publications
Portela, R; Faria, NA; Mwangi, M; Miragaia, M; de Lencastre, H; Tomasz, A; Sobral, RG. 2022. Analysis of a Cell Wall Mutant Highlights Rho-Dependent Genome Amplification Events in Staphylococcus aureus. MICROBIOLOGY SPECTRUM, DOI: 10.1128/spectrum.02483-21
Bárbara VGonçalves; Raquel Portela; Ricardo Lobo; Teresa AFigueiredo; Inês RGrilo; Ana Madalena Ludovice; Hermínia de Lencastre; Jorge SDias; Rita GSobral. 2019. Role of MurT C-Terminal Domain in the Amidation of Staphylococcus aureus Peptidoglycan. ANTIMICROBIAL AGENTS AND CHEMOTHERAPY, DOI: 10.1128/aac.00957-19
Raquel Portela; Catarina RLeal; Pedro LAlmeida; Rita GSobral. 2019. Bacterial cellulose: a versatile biopolymer for wound dressing applications. Microbial Biotechnology, 12(4), DOI: 10.1111/1751-7915.13392
Harrison, Ewan M.; Ba, Xiaoliang; Coll, Francesc; Blane, Beth; Restif, Olivier; Carvell, Henry; Köser, Claudio U.; Jamrozy, Dorota; Reuter, Sandra; Lovering, Andrew; Gleadall, Nicholas; Bellis, Katherine L.; Uhlemann, Anne Catrin; Lowy, Franklin D.; Massey, Ruth C.; Grilo, Inês R.; Sobral, Rita; Larsen, Jesper; Rhod Larsen, Anders; Vingsbo Lundberg, Carina; Parkhill, Julian; Paterson, Gavin K.; Holden, Matthew T.G.; Peacock, Sharon J.; Holmes, Mark A.. 2019. Genomic identification of cryptic susceptibility to penicillins and β-lactamase inhibitors in methicillin-resistant Staphylococcus aureus. Nature Microbiology, DOI: 10.1038/s41564-019-0471-0
Espadinha, D; Sobral, RG; Mendes, CI; Meric, G; Sheppard, SK; Carrico, JA; de Lencastre, H; Miragaia, M. 2019. Distinct Phenotypic and Genomic Signatures Underlie Contrasting Pathogenic Potential of Staphylococcus epidermidis Clonal Lineages. Frontiers in Microbiology, 10, DOI: 10.3389/fmicb.2019.01971
Reichmann, Nathalie T.; Tavares, Andreia C.; Saraiva, Bruno M.; Jousselin, Ambre; Reed, Patricia; Pereira, Ana R.; Monteiro, João M.; Sobral, Rita G.; VanNieuwenhze, Michael S.; Fernandes, Fábio; Pinho, Mariana G.. 2019. SEDS–bPBP pairs direct lateral and septal peptidoglycan synthesis in Staphylococcus aureus. Nature Microbiology, 4, DOI: 10.1038/s41564-019-0437-2
